I originally planned to update this blog every week or so during school, but as soon as the quarter started, things got super busy and it was easy to put this off. Hopefully, I will be better about it this quarter!
To give some background, ever since I started thinking about applying to grad programs, I knew that I wanted to come to my school and program, Biomathematics. I did a lot of research on different aspects of the programs, and even more after I was invited to the interview weekends. I chose this place based on a lot of factors, including academic fit, future goals, advisors, general feel of the program, location, and LGBTQ+ friendliness of the campus.
The program has been wonderful so far and has even surpassed my expectations. It is a pretty tiny program, only 15 grad students total, so the classes are very small and everyone in the program knows each other. Every Thursday, the grad students, some of the students who work for our professors but are from neighboring departments such as Math and Biostatistics, and postdocs all go to a nearby bar, Barney’s, for “pub night”, where they basically drink beer, spill (metaphorical) tea, and relieve stress. In my experience with the students, they have all been incredibly helpful, friendly, and inclusive. I have been careful about sharing personal information with them and thus have only come out to one person in my program so far. I hope that I can make closer friendships with the other students over time.

Grad school classes have been an adjustment in a lot of ways. On one hand, there is a lot more material covered per class – there have been times when the entirety of a math class I had taken in undergrad was covered in just two lectures. Moreover, it is impossible to get all the required background simply from attending class, and it’s necessary to do a lot of extra reading. One thing that has been surprising for me is that in my program’s core courses, as well as the neuroscience course I took, it didn’t seem as difficult as it was in undergrad to get good grades (despite the material being a lot more daunting). I think this is probably because in undergrad, there were more tricks on exams that were designed to weed people out, and now, the focus is on learning, asking questions that may or may not have answers, and being self-motivated to seek extra references for more information, but we aren’t being directly or comparatively evaluated for those things.
Another difference is that there is a lot more emphasis on reading papers and critical thinking, such as proposing potential experiments or critically examining the presentation of data and results in published papers. Some of my core biomathematics courses had homework problems that had no analytic solutions, or that there were multiple possible approaches, and the professors just wanted to see us come up with ideas, defend our assumptions, and solve as far as analytically (or numerically) possible. This is obviously quite different from undergraduate mathematics or chemistry classes, where there are standard solutions to most classical problems either in the back of the book or somewhere on the internet! But I suppose it is moving more reflective of problems in research that have not been previously solved.
I have particularly enjoyed the aspect of courses that involve choosing papers to review for final presentations, and it has allowed me to explore applications of mathematics and computation to neuroscience and has made me more excited about research. When I was in undergrad, although I studied in a theoretical physics group that looked at neuron dynamics, I wasn’t sure if I was doing it only because that was the main opportunity that came my way, but not out of real passion. I think I was too stressed about the prospect of grad school at the time to really develop my passion in research. However, I have always found myself drawn to related topics for class projects and during our department seminars. Biomathematics is a broad field, and I was originally considering exploring the statistical genetics route that is popular in my department, but after starting here, I think that my interests truly lie in neuroscience and mathematical physics, and I am now much more certain in choosing my research focuses and courses.
My department has many course requirements (4 core biomath courses, 2 biomath electives, 6 applied math courses, and 6 biology courses), and as a result, unlike some of the more experimentally focused departments like biology and engineering, they encourage us to focus on coursework and passing the qualifying exams during the first year. We don’t have official research rotations, and we don’t have to decide on an advisor until the end of the second year. However, all of my classmates have started working with potential advisors.
Although I unofficially attended research meetings in fall quarter, this winter quarter was my first official quarter of directed research. At the same time, one of my core courses was taught by my potential advisor (or PI, although my friends who are not in science keep thinking I mean “private investigator” when I use that term). He was an amazing lecturer; he wasn’t the kind of professor who continuously spews information while we furiously try to scribble everything down, but he led us to certain ideas by asking questions. One thing I really like about working with him, both through the course and during the research meetings and updates, is that although his work is clearly mathematically oriented (his background is in particle physics – interestingly, just like my PI in undergrad), unlike a lot of mathematicians and physicists, he has a very conceptual and biologically relevant approach. Some people in our program prefer more mathematical rigor, but for me, it seemed to be a perfect blend.
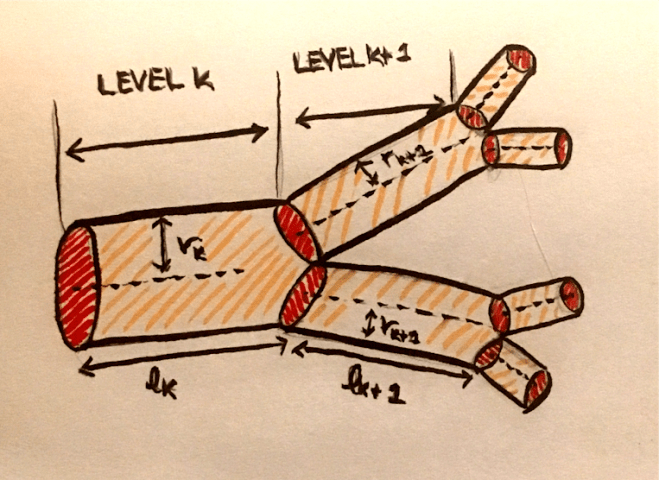
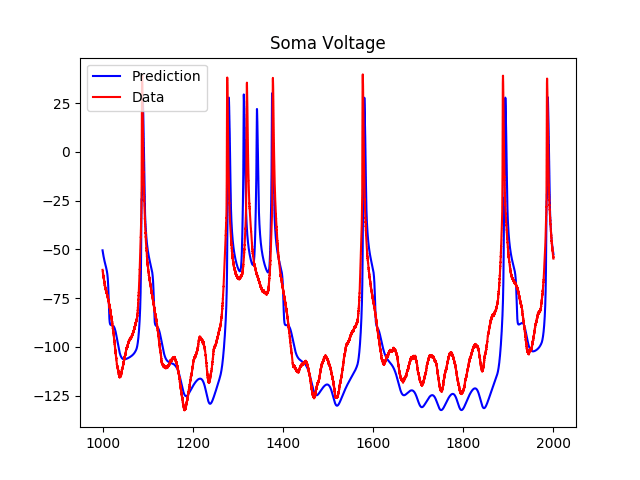
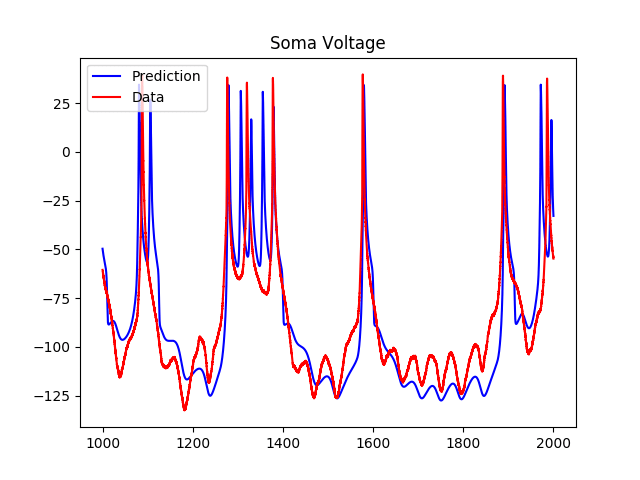
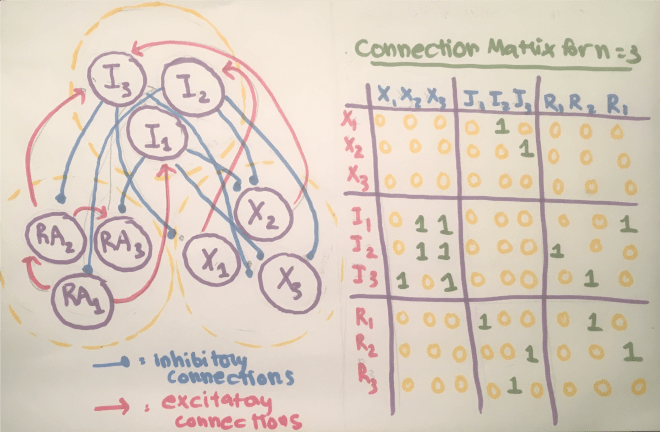
My advisor has done a lot of previous work on cardiovascular networks and the scaling of radius and length of individual vessels across levels of the network. I came to visit him before applying to the program, and when I told him that I was interested in neuroscience, he said that he could imagine the possibility of applying the same methods of analysis to study neuronal networks. Since I came to the group, I have been working on formulating this problem, solving for scaling ratios using Lagrange multipliers (more details about this method in my First Quarter Research Progress post), and analyzing data, both from images and quantitative data from 3D reconstructions of neurons. I have reformulated this problem so that instead of minimizing the power loss due to dissipation, I am minimizing conduction time. For neurons, one of the major evolutionary driving factors is the speed in conducting signals. For example, if you touch something hot like a stove, it would be helpful to have this sensory information relayed as soon as possible so you can pull your hand away before burning it! I have also been reading some papers from the fifties about conduction velocity in neurons and the effects of myelination (fatty layers that provide insulation for nerve fibers) on this speed, and have recently incorporated the degree of myelination as a parameter. I am also looking to modify the space filling constraint to fit neuronal systems, but I am not quite sure how to do this yet. Taking neuroscience courses concurrently with this project is helpful because sometimes I will get random ideas from class that I might be able to translate to math in a way that I can incorporate it into my model. Sometimes, I watch videos of talks by researchers in biology about dendritic morphology and structural neuroscience and feel somewhat overwhelmed, because I am obviously making a lot of simplifying assumptions and not taking into consideration factors such as genetic influences.
Overall, although research is messy and involves a lot of seeking information from various fields, as well as catching up on basic electrodynamics, fluid mechanics, and neuroscience that I never learned in a class, I am enjoying it a lot. This is my first time having my own project, as in undergrad I was for the most part working as a minion, completing menial coding tasks for grad students’ projects. My office mate in my undergrad research group, now a fourth-year grad student in the same group, came to visit me over spring break and told me I seemed a lot more confident than I was last year. Which is strange to me because I feel more overwhelmed and confused the more I learn! I suppose the “confidence” might come from accepting that I don’t know everything, or even a lot, and I’m more comfortable with being uncomfortable, if that makes any sense at all.
As I anticipated, making friends has been quite difficult for me in grad school. It was especially difficult in fall quarter, when I avoided going to LGBTQ+ specific events out of fear of the unknown, mostly, and just went to the weekly department pub nights every now and then, and spent the rest of my time shut up in my own room. My department mates are wonderful and lovely, but aside from the fact that I am not hugely into drinking, the conversations were centered around heteronormative romantic experiences, and I found myself feeling isolated a lot of the time – especially since I’m not out to most of them. When I talked to my mom about it over winter break, she suggested that I add queer org meetings to my schedule rigidly, with the same priority as classes, just so that I could feel more of a sense of community. I decided that this was a good idea, as mental health is an important thing to commit to.
In winter quarter, I regularly attended two queer orgs. One of these is called QSTEM, or Queers in STEM. It was founded by a second year PhD student in Geochemistry who identifies as a gay man. This org is mostly other graduate students, and the vast majority of them are men, which is not entirely unexpected. I have enjoyed participating in social events such as board game nights and ice cream socials. They also have a lot of outreach opportunities, which I hope I have time to get more involved in as my courses finish up and some time is freed up.
The second org I attended was called Queer Girl, and is only open to women and non-binary people. I was the only one there who wasn’t an undergrad, but was a nice social space to discuss things like queer representation in media (or the lack thereof, especially when it comes to women) – it gave me the opportunity to talk about Shay Mitchell in Pretty Little Liars and a random Korean webtoon I found called “Fluttering Feelings.” There’s definitely a lot I could learn from these women, as they would talk about their sexuality openly, which is something I’ve never been comfortable doing. Being around other women like me helped normalize my experiences a little. One of the coordinators of the group was a fellow Asian woman from San Diego (when I went to undergrad), and it was nice to meet someone I could vent to about missing San Diego and people always assuming we’re straight (being Asian/South Asian and having long hair is a surefire way to convince everyone you’re straight).
One of the social events in this club was a trip to Cuties Coffee, a queer owned and themed coffee shop in East Los Angeles that is designed to be a daytime, sober space for queer socialization and an alternative to the gay bars in West Hollywood. I loved visiting this place so much that I have now made it part of my weekend routine – I go there from around noon to four almost every Saturday to either study for classes or work on coding for research. I have included a picture from that day, and used the rainbow pride flag emojis to cover faces for the privacy of the other org members.
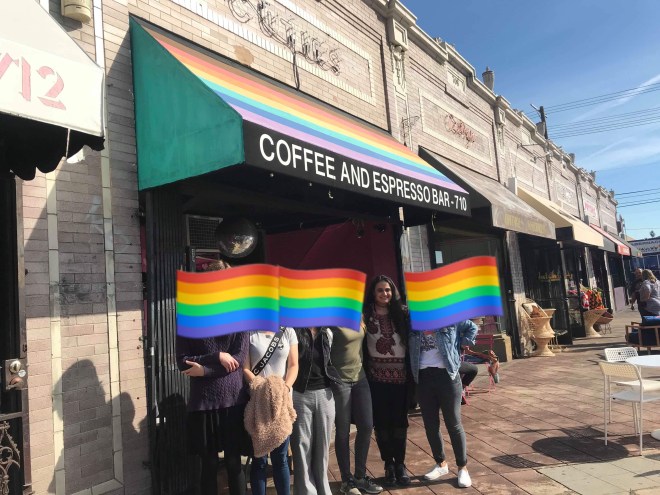

I can’t stress how important it has been for me to have a queer sober space to go to, as I would say I’m pretty far on the introverted side of the spectrum and I never quite feel comfortable meeting new people in bars or nightclubs. (I still mostly keep to myself, drink my coffee/tea, and study during my trips to Cuties, but I hope I will cross the barrier of talking to strangers soon!). At the beginning of winter quarter, I went to West Hollywood a few times to check out the gay bars and nightclubs. Although I love walking on the main strip in West Hollywood, and enjoyed the experience to some extent, it’s not ideal for me because 1) the bars and clubs are largely catered towards gay men – Wednesdays are the only nights specifically for women, and there are no specific clubs for women, and 2) for some reason, being in these spaces where I’m (theoretically) approaching random strangers who are making snap judgments and impressions about me solely based on my physical appearance spiked some of my body insecurities, and to be honest, that’s not a headspace I want (or need) to be in. Right now, the focus for me is on meeting new queer friends and building community, and I’m grateful for these multiple sober spaces I have had access to this quarter.
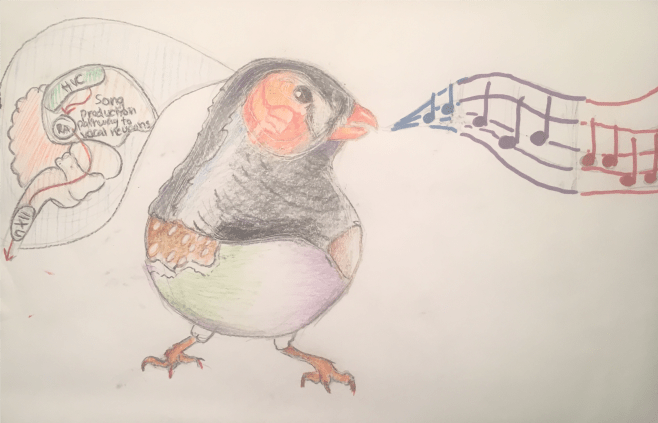
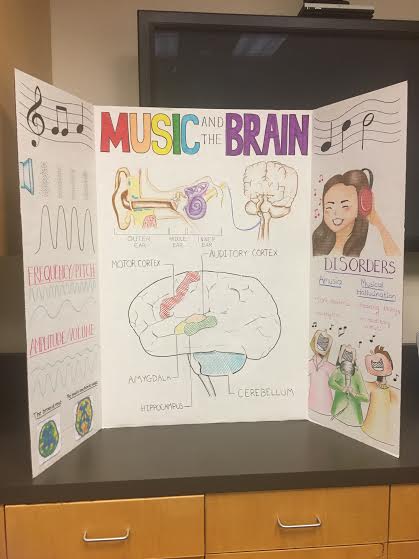
Another extracurricular activity I participated in this winter was a club that does educational outreach in the form of presenting posters about various neuroscience to elementary through high school students to get them excited about learning about the brain. I was part of this Committee called Project Glia, which is responsible for designing and creating posters. I really wanted a way to keep in touch with my art – it can be extremely cathartic and rewarding, and I also want to catch up on the neuroscience background I never had in undergrad for my research, so this was the perfect opportunity for me. I designed this poster for “Music and the Brain”, and I was working with two undergrads who did a lot of the neat typography and shading. The director of Project Glia is a senior undergrad who happens to be taking one of my current graduate neurosciences classes with me, The Biology of Learning and Memory.
One thing I found strange in participating in these activities is that sometimes the undergrads I interact with seem to look up to me in a way, or think that I know things because I am a grad student. One of the students was talking to me the other day about imposter syndrome and comparing yourself to other people, and I ended up saying something along the lines of “Oh, I totally understand that feeling because I used to do that too. But honestly, you can drive yourself crazy comparing yourself to other people – I know because I have done it too, but I realized it was no longer serving me, and I realized I don’t have to be this ‘star student’ to still enjoy what I’m doing.” “That’s SO true,” she had responded sincerely, and meanwhile I was internally panicking during this entire interaction. It was different than listening to a friend, someone who considers me a colleague, and I was suddenly aware of the power dynamic and how much responsibility I had. I think because I’m currently a woman in a grad school program in a related field, some of these women who have goals of grad or med school see me as a sort of safe person to vent to who knows what it’s like to go through this kind of application process and how demoralizing it can be. I was quite nervous about saying the right thing, and having the right mix of relatability and encouragement – all without sounding too preachy or pretentious. When I talked about this later at pub night later with a sixth-year in my program, someone who has significant teaching experience, he reiterated that I have the power to reduce these young women’s imposter syndrome in STEM simply by listening to them and encouraging them. Which is exciting, but also intimidating, because just a year ago, I was that undergrad.
Anyways, that is the (long-winded) gist of the updates of my grad school life over the past quarter. I have some ideas for future, more focused posts, but hope to update more often with these topics as they come up! Until then, I have an exam coming up in my cell neurobiology course, a data analysis assignment, and a research presentation coming up. Wish me luck!