[latexpage]My most important experience in undergrad was working in a group in theoretical physics studying neurons, both on the level of individual neurons and beginning to build simple models for neuronal networks. My group studied a range of nonlinear dynamical systems, and my research focused on dynamics at the molecular level.
When I first began working in the group, my primary prior experience had been undergraduate coursework in chemistry. I had taken only lower-level undergrad courses in math, physics, and to a lesser extent, biology, and my only programming experience was one week of an online course in Python. It definitely didn’t feel like enough at first, and it was an extremely steep learning curve. After my two years of working there, I picked up a lot of skills in programming, learned some basic neuroscience and physics concepts, was able to put the material from my coursework in mathematics, numerical analysis, and programming into practice, and most importantly, learned how to teach myself new material on the fly.
The data I had access to for my research was current and voltage data from current clamp experiments. This means that during the experiment, a current was injected into a cell, and the resulting potential was measured at discrete time intervals of 0.02 milliseconds. Although we only had data from one of the variables, since the dynamical equation of voltage depends on the dynamics of the gating variables and a set of parameters such as the maximal conductances of the ion channels, we can extract this information from the voltage time series. We do this by minimizing a cost function, which has terms for both measurement error and model error. We fix the measurement error and begin with an initial model error, obtaining an initial guess for the minimum, and then me slowly enforce the model constraints until we arrive at a global minimum. We use this state to estimate the most likely values of parameters and time series for the variables.
The first project I worked on was estimating parameter values for induced human neurons. Our experimental collaborators in neuroscience were able to create these cells by converting human skin stem cells to cells with neuronal properties. They were able to obtain current and voltage data through current-clamp experiments. The goal of the project was to estimate parameters for both healthy cells and cells from Alzheimer’s patients. In comparing the results, if we are able to find separation in the parameter space, we might even use this to classify unknown cells based on their current and voltage activity. Moreover, we can learn more about the dynamics and modify our model for induced human neurons as needed.
To test the validity of our estimates, we use the parameter estimates at the end of our time window and use the model to integrate forward the voltage equation, obtaining a time series prediction for voltage. If these predictions match the data closely, we can place more confidence in our estimates.
Using the simple NaKL model, where we were only considering sodium and potassium currents, we got the following results for predictions:
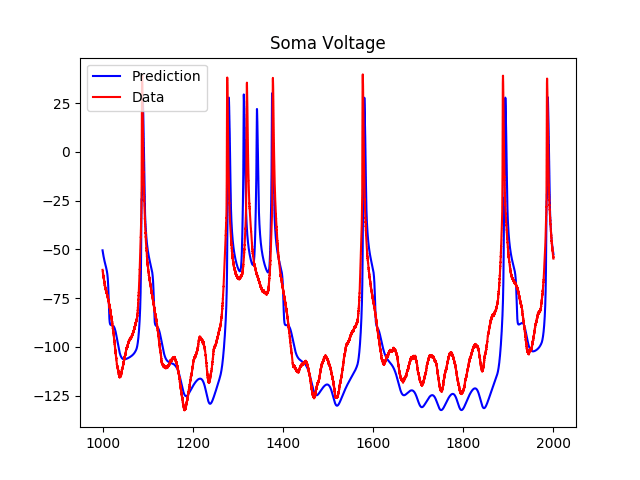
As we can see, although the model predicts the spiking regions well, the subthreshold regions are less accurate. As a result, I tried adding a hyperpolarization-activated inward current to the voltage equation, which added two more variables to the system. The results of the predictions using the estimated parameters were as follows:
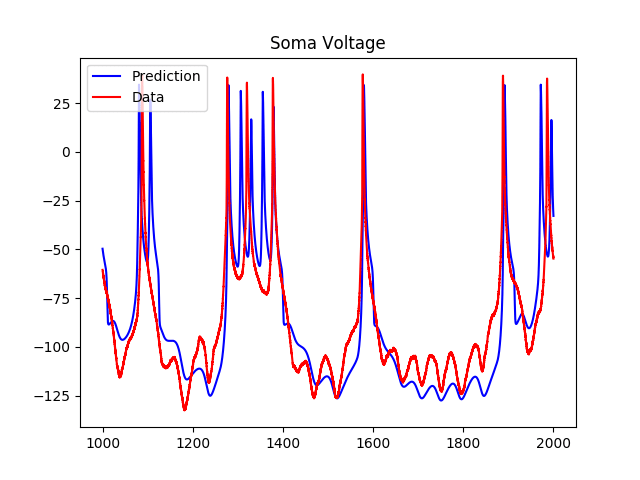
Another project I started working on was modeling the network of neurons in HVC, the premotor nucleus of a songbird called a zebra finch. Songbirds are good models for human language learning because male songbirds spend their youth listening to a tutor, producing syllables and listening to themselves, and eventually establishing a pattern of song syllables unique to themselves.

Within HVC, there are three types of neurons. The $HVC_{Ra}$ neurons lead to the premotor pathway for the song, the $HVC_{X}$ neurons are essential for learning and memory, and the $HVC_I$ neurons have inhibitory connections with the other two types of neurons.
We built a simple model of the connections with the following assumptions, determined from the results of in vivo experiments:
- $HVC_I$ neurons have only inhibitory connections with the others
- $HVC_{RA}$ and $HVC_X$ neurons have only excitatory connections with $HVC_I$ neurons
- $HVC_{RA}$ have a sequence of excitatory connections with each other that store the bird’s own song
- There are no direct connections between $HVC_{RA}$ and $HVC_X$ neurons
- There can be multiple inhibitory connections on a single $HVC_X$ neuron
- The auditory input, which is converted to a current, directly influences all of these neurons to some extent
Below is an illustration of the simplest form of our model, with only three neurons of each type:
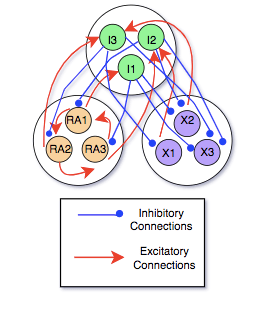
When I was working in the group, we did not yet have experimental data. However, we attempted to create simulated data with pre-determined parameters and use our methods to estimate them. We planned to use the results of these twin experiments to design experiments for our collaborators.
We used song recordings from the lab and extracted pressure wave data from the mp3 files, and then used a transfer function to convert this to a current. Then, we used this current and parameters values we determined, integrating the model’s dynamical equations and obtaining time series data for voltage and the gating variables. In this model, there are nine neurons, and each of these has its own voltage equation and corresponding gating variable equations.
I was only able to complete the twin experiments for this simple model before coming to grad school, but during my time in the group, I developed a script in C that would automatically write the model equations and organize the relevant information into the files we need for data assimilation.
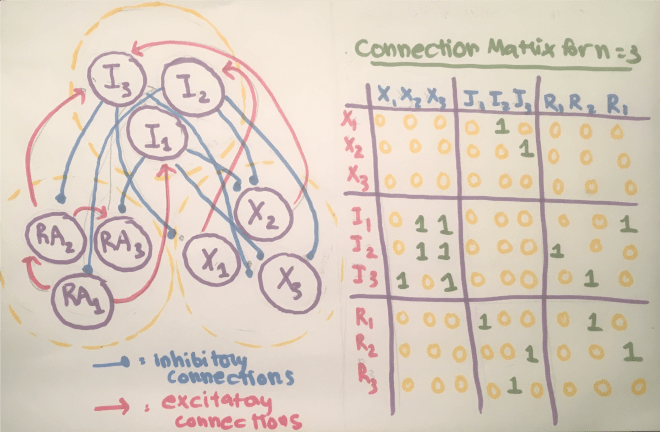
My code makes use of the connection matrix, where the column on the left refers to the presynaptic neuron and the column on the right refers to the postsynaptic neuron, and the synaptic connections strengths are either 0, signaling no connection, or 1, signaling a connection. The code asks the user to manually list the connections using coordinates.
The code can easily be modified for more complex models, such as varying the size of the connection matrix, or varying the strengths of the synaptic connections. When I first wrote the files for data assimilation for this model with a network that has three neurons of each type, it took a couple weeks to complete manually, with some trial and error. My hope that this code will make it more efficient to run twin experiments for larger and more complex models.
I am happy with the research experiences I have had in undergrad, and I feel that it has prepared me to approach independent research here in graduate school. However, our models are very simple and not very biologically realistic. Since my program has a greater emphasis on not only physics, but biological training, I will be able to understand the properties and behavior of neurons at a deeper level, and develop models that are not simply mathematically elegant, but capture the essence of the biology as accurately as possible.
References
Armstrong, E., Abarbanel, H. D. (2016). Model of the songbird nucleus HVC as a network of central pattern generators. Journal of neurophysiology, 116(5), 2405-2419.
Daou, A., Ross, M., Johnson, F., Hyson, R., Bertram, R. (2013). Electrophysiological characterization and computational models of HVC neurons in the zebra finch. Journal of neurophysiology, 110, 1227-1245.
Long, M. A., Jin, D. Z., Fee, M. S. (2010). Support for a synaptic chain model of neuronal sequence generation. Nature, 468(7322), 394.
Mooney, R., Prather, J. F. (2005). The HVC microcircuit: the synaptic basis for interactions between song motor and vocal plasticity pathways. Journal of Neuroscience, 25(8), 1952-1964.
Williams, H. (2004). Birdsong and singing behavior. Annals of the New York Academy of Sciences, 1016(1), 1-30.

