Ever since I found out what qualifying exams were, I was absolutely terrified. I remember being an undergrad listening to the grad students from my research group and my TA sections talking about “that test you have to take after the first couple years where you can be tested on literally anything in your field and if you fail, you get kicked out of grad school lol” and, as someone with low to medium key test anxiety, it sounded like my personal kind of hell. Even after going through the grad school application process, my entire future rested on a few hours and a few pieces of paper?
Our written quals are subject-based. We have five core courses: Deterministic Models in Biology, Modeling in Biology: Structure, Function, and Evolution, Stochastic Modeling in Biology, Biomedical Data Analysis, and Computational Algorithms. The qualifying exams for those subjects are offered at the end of August each year. Each subject exam can be assigned a PhD pass, a Masters pass (slightly lower level), or no pass. In order to pass the overall comprehensive exams and remain in the program, a student must get at least three PhD level passes and one Masters level pass. Each student gets two tries to get the required number of passes.
It might seem like these qualifying exams are just like final exams, since, after all, they are single exams self-contained in just five 10 week courses, right? Wrong! What I quickly learned when I entered grad school was how much all of these courses built on years and years of knowledge from high school and undergrad mathematics, how much of this knowledge was assumed background knowledge that was required in order to even begin to comprehend any of the lectures. I realized how kind my undergrad professors and TAs had been in taking the time to rehash material from basic algebra 2, trigonometry, and differential equations in office hours in order to help us understand more difficult material. I missed the warm embrace of assumed ignorance, as my graduate school professors were surprised, disappointed, and in some cases even mortally offended if students showed the slightest sign of rustiness in material we should have learned in our undergrad probability theory courses, in our numerical linear algebra courses, and in our complex analysis courses. It was intimidating, to say the very least, and I certainly did not pick up all of the material from the lectures the first time around. Aside from reviewing all my undergrad course notes and textbooks and completing all the core course problem sets on time, there was so much to do and so much to learn during the quarter with research, preparing for group meetings, and neuroscience electives. Throughout the year, the prospect of qualifying exams seemed to be looming over my head. To the put it in the most graceful and delicate way possible, I was terrified because I didn’t know s***.
During the first part of the summer, before my San Diego Pride trip, I spent my days in the lab, partly working on research and partly reviewing and rewriting all my notes from the lectures and the textbooks. After the trip, after recovering for a few days, I collected myself (physically and emotionally) and collected all the books and notes from undergrad that I thought would be useful in order to decode the notes that I had spent the first half of the summer writing. First, I went through all the problem sets that I had already done during the quarter. Looking at the solutions I had written up (most of which I had forgotten by this point), I tried to recall the theorems from undergrad courses I had used, and the corresponding textbooks that would have more detailed information I could review. After finding these textbooks all around the various bookshelves in the house, I went through the sections I thought would be useful. Below is a stack of the textbooks that I used during this process.

For me personally, I found that the most gaps in my knowledge were in probability and linear algebra, as my Stochastic Modeling and Computational Algorithms classes (both taught by the same professor) took a lot of the theorems and proofs I learned in those courses for granted.
Something I really came to appreciate through studying for these exams was the sheer intellectual brilliance of my professor who taught my Stochastic Modeling and Computations Algorithms courses. He had written textbooks for these courses, and I am ashamed to admit that during the classes, I had skipped over many of the proofs and examples in the book. A fourth-year student in my department and in my lab, one of the few people who entered my program with more of a biology background than a math background, shared some advice on passing this professor’s exams, for someone with less confidence in their mathematical abilities. “Read all the examples and proofs in the textbook. Make sure you can understand how he got to the conclusions. His books are very dense and compact and he skips a lot of steps. Make sure you know how to fill in the gaps.” This seemed like a daunting task, but this older student (bless his soul) also provided me with a 75 page stack of his notes on the textbook examples from the time when he was studying for quals three years ago, where he filled in the gaps, and I could use them as a reference in case I got stuck. For example, my professor used things like binomial theorem and Taylor expansion approximations to condense a lot of the equations, things that weren’t immediately obvious upon first glance. It was a daunting task, and I didn’t get through the textbooks cover to cover. But I got through a significant portion of the chapters that were more emphasized in the courses, and in the end, I felt like a stronger applied mathematician. My eye had gotten better at recognizing when to use these little tricks to simplify expressions and approximate.
One of the courses, the Structure, Function, and Evolution course, was taught by my own PI, which meant it would be important for me to pass this particular subject because I do want to remain in his lab. The interesting thing about his course is that it was not mathematically the most challenging, although there were some complicated PDEs there when we started talking about diffusion and population genetics. During the classes, when we had problem sets due, his office hours would be completely full with students from the course probing him what exactly he was trying to ask with the questions and trying to decode his convoluted wording. According to the older students in our program, the main difficulty about his exam was interpreting the questions. After looking at some of the past exams, I noticed some general themes, as he tended to ask questions that bridged concepts we learned earlier in the class, relating to network theory and geometry, and later concepts in population genetics.
Biomedical Data Analysis was one that I felt least prepared for, as during the class, we had focused a lot on using R to extract statistical parameters from datasets and fit models to the data, but not much on deriving statistical results. Particularly for me, since I did not have much of a statistics background in undergrad, I felt even more overwhelmed by the unfamiliar vocabulary that my professor assumed we had learned in kindergarten. I studied for this one by making a lot of use of statistics videos on YouTube, which proved to be more useful in aiding my conceptual understanding than our textbook. In addition, our professor was kind enough to host a review session at the beginning of August, which clarified some of the confusion I had.
The older students in our department had told us that very few students in our department had obtained PhD level passes on one of the exams, the Deterministic Models in Biology Course. It was taught by a notoriously tough professor with a background in Physics and a joint appointment in the Mathematics department, and many of the homework problems he had assigned didn’t even have analytical solutions. Since my main goal was to stay in the program and I only had a couple months to study for these exams, I followed the advice of the older students and spent the majority of my time on the other subjects, leaving only a couple of days to study for that one.
Before August, I had spent most of my time studying at home, since I did not want to lug all my undergrad textbooks around on campus. However, I believe that in preparation for exams as intense as these ones, it can be very helpful to study with others to get some feedback and test understanding. During August, I spent a lot of my time studying with my classmate Janet* (name has been changed for privacy). She is the only other PhD student in my year and already has a medical degree and had studied in China, where early math education was much more advanced, so I definitely worried that our study groups would end up being her incredible brain carrying a lot of my dead weight. I had declared my major in math late, and didn’t know what proof by induction was until the latter half of my second to last year of college. Meanwhile, she already knew how to apply proof by induction while crawling out of the womb (okay, this *might* be a *slight* exaggeration, but truly not that far off). However, I think we got a really good, productive, mutually beneficial flow going when we were studying together in August.
At first, I spent the whole day studying in the office, but after a while, I realized that being around the older students stressed me out more than it helped. One of my pet peeves was when they would try to quiz me on random facts from some of the courses, shouting at me things like “Hey, quick, under what conditions can you add the powers when multiplying matrix exponential? When the power matrices commute, duh! Those are easy points you’re missing!” Of course, I didn’t know how to conjure these facts on the spot, but I felt that I did know more than it seemed from my blank looks, because after thinking about it for a moment, I could even conjure a proof for that fact. Although these students were well-intentioned, I knew what worked best for me, and it was not being holed up in the office all day, subject to this stressful banter that left me feeling discouraged about my prospects for the exams.
This roadblock turned out to be a blessing in disguise, because I soon fell into the easy routine of going to the campus at around 7 am, spending the day reviewing past exams in the Biomedical Library until 3 in the afternoon. From around 3-5 pm, I would go up to the office to discuss these exams with Janet. I found that though she helped a lot with the more probability theory related problems, I was also able to help her a lot with my PI’s convoluted wording in his past exams due to the language barrier. Plus, I felt that after working for my PI for almost a year, I got a sense of how his brain worked and the kinds of questions he was asking. I was glad that I could contribute to these study sessions as well as gain from them. I think this process of studying improved my work ethic and anxiety management, forced me to review individual undergrad courses and bring them together in ways that I didn’t know existed, and improved my confidence in problem-solving. Something I think about a lot is how in undergrad, I took a variety of applied math courses, but learned about mathematics mostly from a theoretical perspective without truly understanding how to apply what I learned to research. I think that the process of studying for these five courses in-depth helped me understand not only what mathematical tools are available, but how to use them in real biomedical problems and why they’re important.
Finally, at the end of August, the exams began. We had three, spaced out days of exams. The first two days, we had two exams each. The first day was Stochastic Modeling in the morning and Computational Algorithms in the afternoon. The second day was Structure, Function, and Evolution in the morning and Biomedical Data Analysis in the afternoon. The last day was just Deterministic Models in the afternoon. For each session, we got a 30 minute reading period, where we could read the exam questions and ask the professors for any clarification about the wording of the questions. Then, the three of us were split off into three separate rooms on the floor. I was assigned the classroom where most of our courses had occurred, which was encouraging because I had read research claiming that recall of material during exams can be enhanced if the exam takes place in the same room where learning occurred (to be fair, though, most of my learning had occurred during the summer at home, in the Biomedical Library, and in the office rather than the classroom). We were allowed to eat and drink during the exams, and the older students were very nice and brought us chocolates and water the day of our first exam.
I will admit that after every single exam, I felt terrible and slightly violated, although none more than the last exam, for which I didn’t even finish half of the questions. The good thing is that for a lot of the exams, it was not necessary to answer all of the questions to completion get a PhD pass; it was more important to show how we are thinking – something I had been trained to do since my elementary school math (“show your work!” is permanently etched in my brain).
After the exams, I took a yoga class with one of my college friends, ran a lot, swam a lot, bought all my textbooks, binders, notebook paper, and replenished pencils for my fall classes, worked on my poster for a quantitative and computational biology retreat where I’m presenting at the end of September, and went to a Diversity in STEM Conference in Irvine where I got to catch up with a friend who is a PhD student there. It was busy, but I needed to keep busy so I wouldn’t keep thinking about my anxiety about the results.
A week later, much earlier than I was expecting, I got the results: I got a PhD level pass in all the exams except Deterministic Models in Biology – I got no pass in that subject. I learned that Janet also got no pass in that exam, and since she’s one of the smartest people I know, in a twisted way, it made me feel a little validated that it’s not like only dumb people get “no pass” or something! (I’m saying this slightly in jest, as I do recognize it as a toxic thought, but it will take some more time to train myself to not have these thoughts.) I will be taking a 2 quarter sequence in Mathematical Physics in the Physics department this coming year, so hopefully, I will fill some of the gaps in my knowledge on that side of Biomathematics. Overall, I’m pretty happy with my results in all the other classes, thrilled that I get to stay in the program and continue working on the project I’ve been working on, looking forward to my last year of courses – all very interesting elective courses I chose because of their relevance to my research – and very much looking forward to meeting all the new grad students in my department (there are five, and mostly other women, by the way, which makes me happy).
This coming Monday is our Department Orientation for the new students, and at noon, there is a potluck where everyone from the department meets the new students. I remember last year, when I was a first-year coming into the department, the Vice Chair announced that both the two second years had passed their qualifying exams. It might seem silly, but during my pre-exam anxiety and habitual catastrophic thinking, I remember thinking about how that if I didn’t pass my quals, it would be announced to all the new entering students, and then I would have to go through this same process again with the first-years next summer. I’m very relieved this will not be the case.
Overall, although I know I will probably forget most of what I learned during this summer, it was helpful for me to have a broad idea about the vast breadth of tools in applied mathematics – knowledge that I will be building on this year in my applied math and physics elective courses. I might not remember the details of how to solve every type of problem by hand, but generally knowing what kind of tools are available, I believe, will make me more informed and better able to come up with ideas to tackle new problems in my research in the years to come. The details are things that I can learn on the fly, as needed.






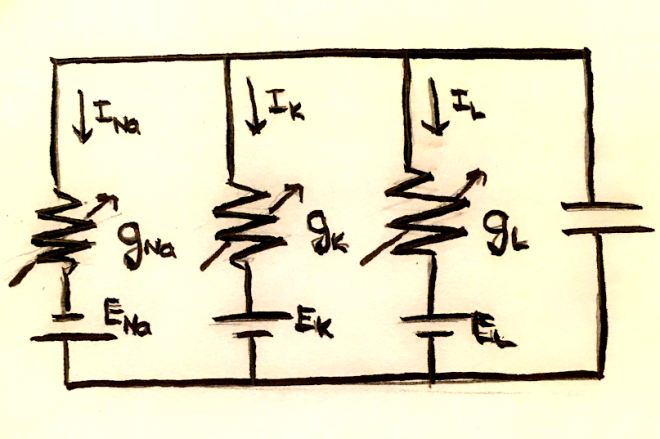 Because of charge conservation, the sum of the currents across the capacitor and each of the resistors must be 0. In mathematical terms, this is $C_m \frac{dV_m}{dt} = -\sum_i I_i$.
Because of charge conservation, the sum of the currents across the capacitor and each of the resistors must be 0. In mathematical terms, this is $C_m \frac{dV_m}{dt} = -\sum_i I_i$.